Note
Go to the end to download the full example code.
Contrast Metric for Calibrated Phantoms#
This example demonstrates contrast measurement using a calibrated CIRS 040GSE phantom, which is helpful for validating beamforming algorithms against known specifications.
Mathematical Definition#
The contrast between two regions (denoted inside \(i\) and outside \(o\)) is defined as:
where:
are, respectively, the mean signal power inside and outside the lesion, where \(s\) denotes the signal. Contrast can take any positive real value, and \(C \to \infty\) as \(\mu_o \to 0\).
Contrast is often expressed in decibels as:
When to use contrast:
Calibrated phantoms with known lesion contrast levels
Linear imaging where intensity directly relates to scattering strength
Validation against manufacturer specifications
When to use alternatives:
gcnr()for nonlinear imaging methods (e.g., coherence-based beamforming)cnr()when accounting for noise statistics is importantDynamic range compression affects traditional contrast measurements
This example uses the contrast metric as described in [1] and demonstrates analysis using the PICMUS dataset [2].
References#
Example#
import matplotlib.colors as mcolors
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from matplotlib.patches import Patch
from ultrasound_metrics.data import db_zero
from ultrasound_metrics.data.uff import load_uff_dataset
from ultrasound_metrics.metrics.contrast import contrast
from ultrasound_metrics.roi.masks import build_mask
Load the ultrasound dataset#
# Load the PICMUS contrast/speckle dataset which contains lesions with known contrast levels
dataset_name = "picmus_contrast_experiment"
image, scan = load_uff_dataset(dataset_name)
image = image.T
extent = (scan.x_axis.min(), scan.x_axis.max(), scan.z_axis.max(), scan.z_axis.min())
Downloading PICMUS_experiment_contrast_speckle.uff: 0%| | 0.00/146M [00:00<?, ?B/s]
Downloading PICMUS_experiment_contrast_speckle.uff: 1%| | 1.05M/146M [00:00<00:29, 4.86MB/s]
Downloading PICMUS_experiment_contrast_speckle.uff: 5%|▌ | 7.34M/146M [00:00<00:05, 27.3MB/s]
Downloading PICMUS_experiment_contrast_speckle.uff: 12%|█▏ | 16.8M/146M [00:00<00:02, 50.6MB/s]
Downloading PICMUS_experiment_contrast_speckle.uff: 18%|█▊ | 26.2M/146M [00:00<00:01, 63.0MB/s]
Downloading PICMUS_experiment_contrast_speckle.uff: 25%|██▌ | 36.7M/146M [00:00<00:01, 74.8MB/s]
Downloading PICMUS_experiment_contrast_speckle.uff: 32%|███▏ | 47.2M/146M [00:00<00:01, 78.5MB/s]
Downloading PICMUS_experiment_contrast_speckle.uff: 40%|███▉ | 57.7M/146M [00:00<00:01, 85.6MB/s]
Downloading PICMUS_experiment_contrast_speckle.uff: 46%|████▌ | 67.1M/146M [00:00<00:00, 84.4MB/s]
Downloading PICMUS_experiment_contrast_speckle.uff: 53%|█████▎ | 76.5M/146M [00:01<00:00, 86.8MB/s]
Downloading PICMUS_experiment_contrast_speckle.uff: 59%|█████▉ | 86.0M/146M [00:01<00:00, 86.3MB/s]
Downloading PICMUS_experiment_contrast_speckle.uff: 66%|██████▋ | 96.5M/146M [00:01<00:00, 91.4MB/s]
Downloading PICMUS_experiment_contrast_speckle.uff: 73%|███████▎ | 106M/146M [00:01<00:00, 88.4MB/s]
Downloading PICMUS_experiment_contrast_speckle.uff: 79%|███████▉ | 115M/146M [00:01<00:00, 86.4MB/s]
Downloading PICMUS_experiment_contrast_speckle.uff: 86%|████████▌ | 125M/146M [00:01<00:00, 81.7MB/s]
Downloading PICMUS_experiment_contrast_speckle.uff: 92%|█████████▏| 133M/146M [00:01<00:00, 79.9MB/s]
Downloading PICMUS_experiment_contrast_speckle.uff: 97%|█████████▋| 142M/146M [00:01<00:00, 79.0MB/s]
Downloading PICMUS_experiment_contrast_speckle.uff: 100%|██████████| 146M/146M [00:01<00:00, 75.8MB/s]
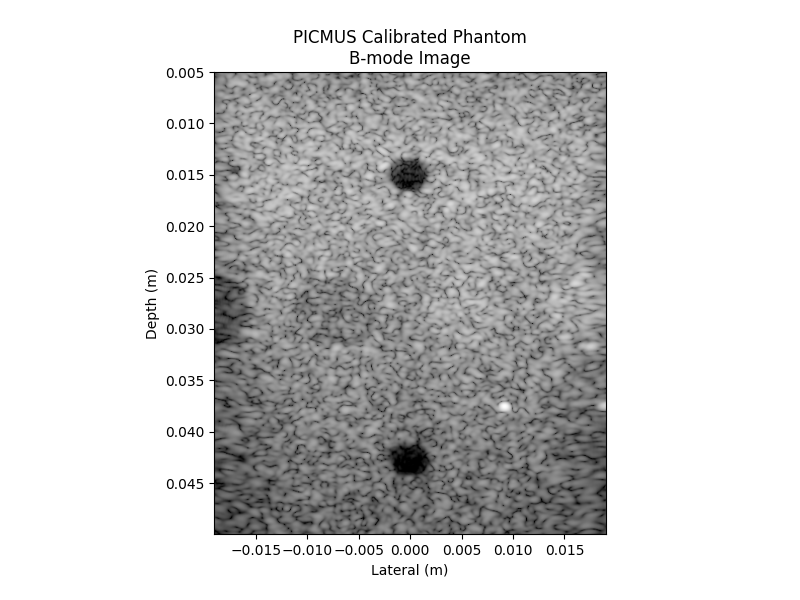
Visualize the B-mode image#
image_db = db_zero(image)
fig, ax = plt.subplots(figsize=(8, 6))
ax.imshow(image_db, cmap="gray", extent=extent, vmin=-60, vmax=0)
ax.set_title("PICMUS Calibrated Phantom\nB-mode Image")
ax.set_xlabel("Lateral (m)")
_ = ax.set_ylabel("Depth (m)")

Define known lesion locations based on PICMUS phantom geometry#
Define lesion locations and compute contrast PICMUS phantom specifications: 8mm diameter (4mm radius) lesions at -3dB and +3dB contrast
lesion_specs = [
{"name": "-3dB lesion", "center": (-0.007, 0.028), "expected_contrast_db": -3},
{"name": "+3dB lesion", "center": (0.0055, 0.028), "expected_contrast_db": +3},
]
# https://www.cirsinc.com/wp-content/uploads/2019/04/040GSE-DS-120418.pdf
lesion_radius = 0.004
# include the background region, but don't include other lesions
background_radius = lesion_radius * 2
Compute contrast for each lesion ROI#
results_data = []
for spec in lesion_specs:
center = spec["center"]
mask_lesion = build_mask(center, lesion_radius, scan.x_axis, scan.z_axis, "circle")
mask_background_and_lesion = build_mask(center, background_radius, scan.x_axis, scan.z_axis, "circle")
mask_background = mask_background_and_lesion & ~mask_lesion
# Compute contrast between lesion and background regions
measured_db = contrast(image[mask_lesion], image[mask_background], db=True)
error_db = measured_db - spec["expected_contrast_db"]
results_data.append({
"Lesion": spec["name"],
"Expected (dB)": spec["expected_contrast_db"],
"Measured (dB)": measured_db,
"Error (dB)": error_db,
"mask_lesion": mask_lesion,
"mask_background": mask_background,
})
Display results table#
results_df = pd.DataFrame(results_data)
print("=== Contrast Measurement Results ===")
print(results_df[["Lesion", "Expected (dB)", "Measured (dB)", "Error (dB)"]].round(2))
# Summary statistics
errors = results_df["Error (dB)"].values
print(f"\nMean absolute error: {np.mean(np.abs(errors)):.2f} dB")
print(f"RMS error: {np.sqrt(np.mean(errors**2)):.2f} dB")
=== Contrast Measurement Results ===
Lesion Expected (dB) Measured (dB) Error (dB)
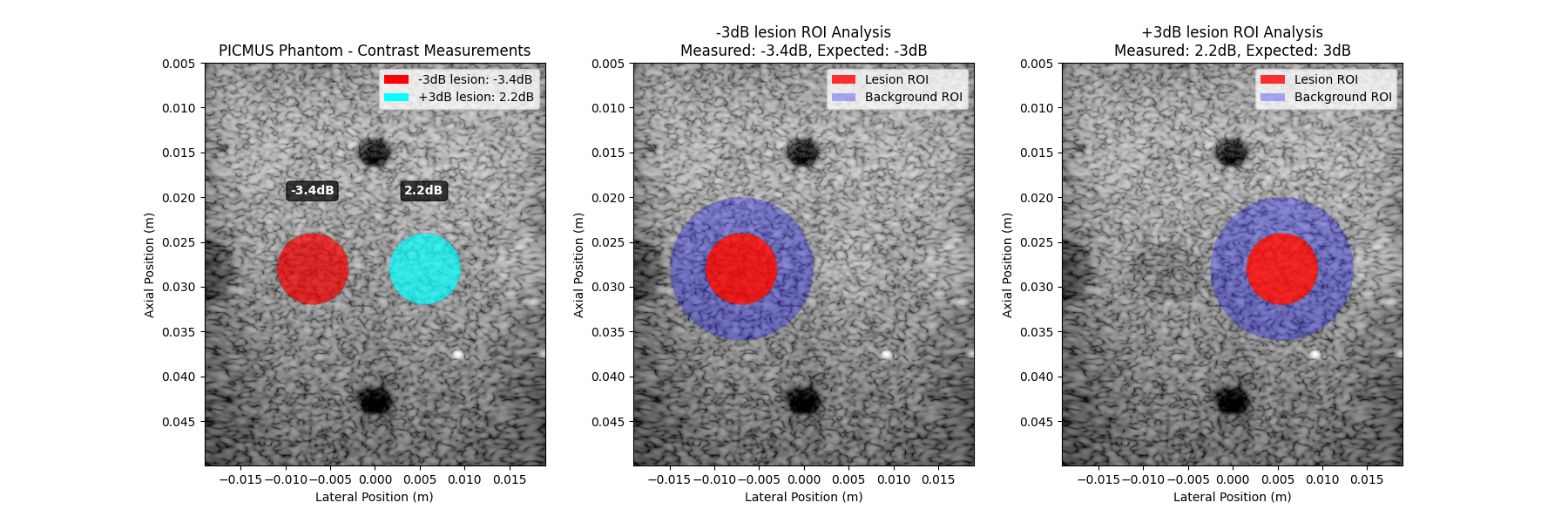
0 -3dB lesion -3 -3.44 -0.44
1 +3dB lesion 3 2.20 -0.80
Mean absolute error: 0.62 dB
RMS error: 0.65 dB
The contrast measurements are close but slightly more negative than expected. The phantom might not be perfectly calibrated.
Visualize ROIs and measurements#
fig, axes = plt.subplots(1, 1 + len(results_df), figsize=(18, 6))
# First subplot: Overview with all lesions
ax1 = axes[0]
ax1.imshow(image_db, cmap="gray", extent=extent, vmin=-60, vmax=0)
colors = ["red", "cyan"]
legend_elements = []
for i, (_, row) in enumerate(results_df.iterrows()):
# Overlay lesion ROI
lesion_overlay = np.ma.masked_where(~row["mask_lesion"], np.ones_like(image_db))
ax1.imshow(lesion_overlay, alpha=0.7, extent=extent, cmap=mcolors.ListedColormap([colors[i]]))
# Add measurement labels
center = lesion_specs[i]["center"]
ax1.text(
center[0],
center[1] - 0.008,
f"{row['Measured (dB)']:.1f}dB",
ha="center",
va="bottom",
color="white",
fontweight="bold",
bbox={"boxstyle": "round,pad=0.3", "facecolor": "black", "alpha": 0.7},
)
legend_elements.append(Patch(facecolor=colors[i], label=f"{row['Lesion']}: {row['Measured (dB)']:.1f}dB"))
ax1.set_title("PICMUS Phantom - Contrast Measurements")
ax1.set_xlabel("Lateral Position (m)")
ax1.set_ylabel("Axial Position (m)")
ax1.legend(handles=legend_elements, loc="upper right")
# Individual lesion subplots: Show each lesion with its background ROI
for i, (_, row) in enumerate(results_df.iterrows()):
ax = axes[i + 1]
ax.imshow(image_db, cmap="gray", extent=extent, vmin=-60, vmax=0)
# Show lesion (red) and background (blue) ROIs
lesion_overlay = np.ma.masked_where(~row["mask_lesion"], np.ones_like(image_db))
bg_overlay = np.ma.masked_where(~row["mask_background"], np.ones_like(image_db))
ax.imshow(lesion_overlay, alpha=0.8, extent=extent, cmap=mcolors.ListedColormap(["red"]))
ax.imshow(bg_overlay, alpha=0.3, extent=extent, cmap=mcolors.ListedColormap(["blue"]))
ax.set_title(
f"{row['Lesion']} ROI Analysis\nMeasured: {row['Measured (dB)']:.1f}dB, Expected: {row['Expected (dB)']}dB"
)
ax.set_xlabel("Lateral Position (m)")
ax.set_ylabel("Axial Position (m)")
roi_legend = [
Patch(facecolor="red", alpha=0.8, label="Lesion ROI"),
Patch(facecolor="blue", alpha=0.3, label="Background ROI"),
]
ax.legend(handles=roi_legend, loc="upper right")
Show all figures (if run as a script)
plt.show()
Total running time of the script: (0 minutes 3.601 seconds)