Note
Go to the end to download the full example code.
Tenengrad Sharpness Metric#
This example uses a PICMUS dataset to demonstrate how to:
Compute and visualize the Tenengrad sharpness metric [1]
Compare sharpness between datasets with different characteristics
Show how the metric responds to different ROI selections
[2] uses the Tenengrad sharpness metric to compare the lateral focusing quality across different speed-of-sound profiles.
References#
Example#
1. Import required modules and functions#
import matplotlib.pyplot as plt
import numpy as np
from ultrasound_metrics.data.uff import load_uff_dataset
from ultrasound_metrics.data.visualize_bmode import db_zero
from ultrasound_metrics.metrics.sharpness import tenengrad
from ultrasound_metrics.roi.masks import build_mask
2. Load PICMUS datasets#
image_resolution, scan_resolution = load_uff_dataset("picmus_resolution_experiment")
image_resolution = image_resolution.T
image_contrast, scan_contrast = load_uff_dataset("picmus_contrast_experiment")
image_contrast = image_contrast.T
print(f"Resolution/distortion dataset image shape: {image_resolution.shape}")
print(f"Contrast/speckle dataset image shape: {image_contrast.shape}")
Resolution/distortion dataset image shape: (609, 387)
Contrast/speckle dataset image shape: (609, 387)
3. Calculate edge-enhanced image#
This is an intermediate step for Tenengrad sharpness.
sharpness_img_resolution = tenengrad(image_resolution, reduce_sum=False)
print("sharpness image shape:", sharpness_img_resolution.shape, "(resolution/distortion dataset)")
sharpness_img_contrast = tenengrad(image_contrast, reduce_sum=False)
print("sharpness image shape:", sharpness_img_contrast.shape, "(contrast/speckle dataset)")
sharpness image shape: (609, 387) (resolution/distortion dataset)
sharpness image shape: (609, 387) (contrast/speckle dataset)
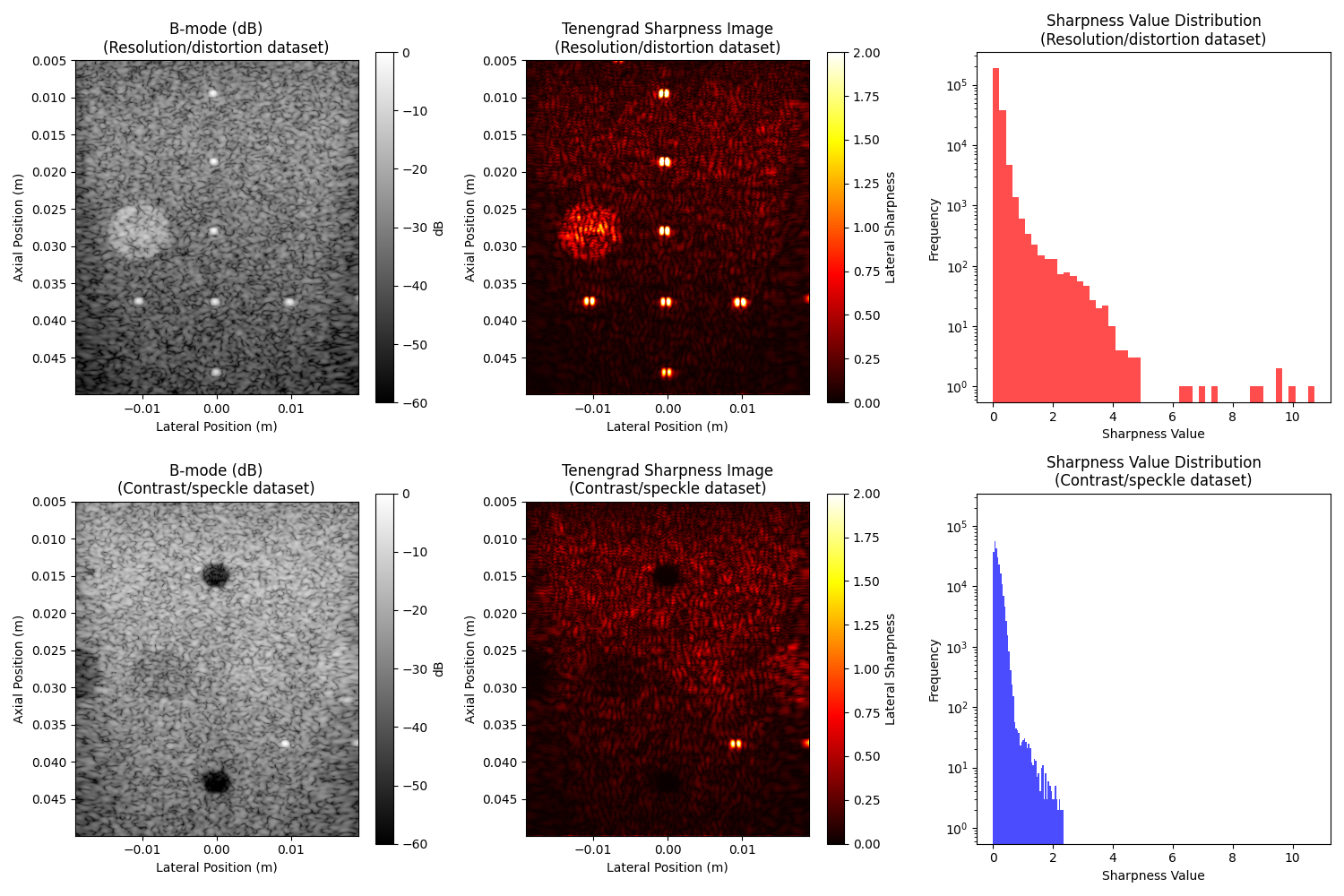
4. Visualize the original images and their sharpness maps#
fig, axes = plt.subplots(2, 3, figsize=(15, 10), gridspec_kw={"width_ratios": [1, 1, 1]})
# Share x and y axes for the histogram plots (right column)
axes[0, 2].sharex(axes[1, 2])
axes[0, 2].sharey(axes[1, 2])
# Get coordinate arrays for proper axis labeling
x_coords_res = scan_resolution.x_axis
z_coords_res = scan_resolution.z_axis
x_coords_con = scan_contrast.x_axis
z_coords_con = scan_contrast.z_axis
# Define common visualization parameters
vmin = -60 # dB range for B-mode images
vmax = 0
sharpness_vmin = 0
sharpness_vmax = 2
# Row 1: Resolution/distortion dataset
extent_res = [x_coords_res.min(), x_coords_res.max(), z_coords_res.max(), z_coords_res.min()]
im1 = axes[0, 0].imshow(db_zero(image_resolution), cmap="gray", vmin=vmin, vmax=vmax, extent=extent_res)
axes[0, 0].set_title("B-mode (dB)\n(Resolution/distortion dataset)")
axes[0, 0].set_xlabel("Lateral Position (m)")
axes[0, 0].set_ylabel("Axial Position (m)")
im2 = axes[0, 1].imshow(
sharpness_img_resolution, cmap="hot", extent=extent_res, vmin=sharpness_vmin, vmax=sharpness_vmax
)
axes[0, 1].set_title("Tenengrad Sharpness Image\n(Resolution/distortion dataset)")
axes[0, 1].set_xlabel("Lateral Position (m)")
axes[0, 1].set_ylabel("Axial Position (m)")
# Histogram of sharpness values for resolution/distortion dataset
axes[0, 2].hist(sharpness_img_resolution.flatten(), bins=50, alpha=0.7, color="red")
axes[0, 2].set_title("Sharpness Value Distribution\n(Resolution/distortion dataset)")
axes[0, 2].set_xlabel("Sharpness Value")
axes[0, 2].set_ylabel("Frequency")
axes[0, 2].set_yscale("log")
# Row 2: Contrast/speckle dataset
extent_con = [x_coords_con.min(), x_coords_con.max(), z_coords_con.max(), z_coords_con.min()]
im3 = axes[1, 0].imshow(db_zero(image_contrast), cmap="gray", vmin=vmin, vmax=vmax, extent=extent_con)
axes[1, 0].set_title("B-mode (dB)\n(Contrast/speckle dataset)")
axes[1, 0].set_xlabel("Lateral Position (m)")
axes[1, 0].set_ylabel("Axial Position (m)")
im4 = axes[1, 1].imshow(sharpness_img_contrast, cmap="hot", extent=extent_con, vmin=sharpness_vmin, vmax=sharpness_vmax)
axes[1, 1].set_title("Tenengrad Sharpness Image\n(Contrast/speckle dataset)")
axes[1, 1].set_xlabel("Lateral Position (m)")
axes[1, 1].set_ylabel("Axial Position (m)")
# Histogram of sharpness values for contrast experiment
axes[1, 2].hist(sharpness_img_contrast.flatten(), bins=50, alpha=0.7, color="blue")
axes[1, 2].set_title("Sharpness Value Distribution\n(Contrast/speckle dataset)")
axes[1, 2].set_xlabel("Sharpness Value")
axes[1, 2].set_ylabel("Frequency")
axes[1, 2].set_yscale("log")
# Add colorbars
plt.colorbar(im1, ax=axes[0, 0], label="dB")
plt.colorbar(im3, ax=axes[1, 0], label="dB")
plt.colorbar(im2, ax=axes[0, 1], label="Lateral Sharpness")
plt.colorbar(im4, ax=axes[1, 1], label="Lateral Sharpness")
plt.tight_layout()
plt.show()
5. Calculate overall sharpness metrics for both datasets#
print("\n=== Overall Tenengrad Sharpness Comparison ===")
# Calculate total sharpness (sum) and normalized sharpness (mean)
sharpness_resolution = tenengrad(image_resolution, normalize=False)
sharpness_normalized_resolution = tenengrad(image_resolution, normalize=True)
sharpness_contrast = tenengrad(image_contrast, normalize=False)
sharpness_normalized_contrast = tenengrad(image_contrast, normalize=True)
print("Resolution experiment:")
print(f" Total sharpness: {float(sharpness_resolution):.1f}")
print(f" Normalized sharpness: {float(sharpness_normalized_resolution):.4f}")
print("\nContrast experiment:")
print(f" Total sharpness: {float(sharpness_contrast):.1f}")
print(f" Normalized sharpness: {float(sharpness_normalized_contrast):.4f}")
# Calculate ratio
ratio = float(sharpness_normalized_resolution) / float(sharpness_normalized_contrast)
print(f"\nSharpness ratio (Resolution/Contrast): {ratio:.2f}")
if ratio > 1.2:
print("✅ Resolution/distortion dataset shows higher sharpness as expected!")
print(" Point targets create sharper edges than diffuse lesions.")
elif ratio > 0.8:
print("️Similar overall sharpness between datasets.")
else:
print("⚠️ Contrast/speckle dataset shows higher sharpness.")
=== Overall Tenengrad Sharpness Comparison ===
Resolution experiment:
Total sharpness: 35788.9
Normalized sharpness: 0.1519
Contrast experiment:
Total sharpness: 35172.2
Normalized sharpness: 0.1492
Sharpness ratio (Resolution/Contrast): 1.02
️Similar overall sharpness between datasets.
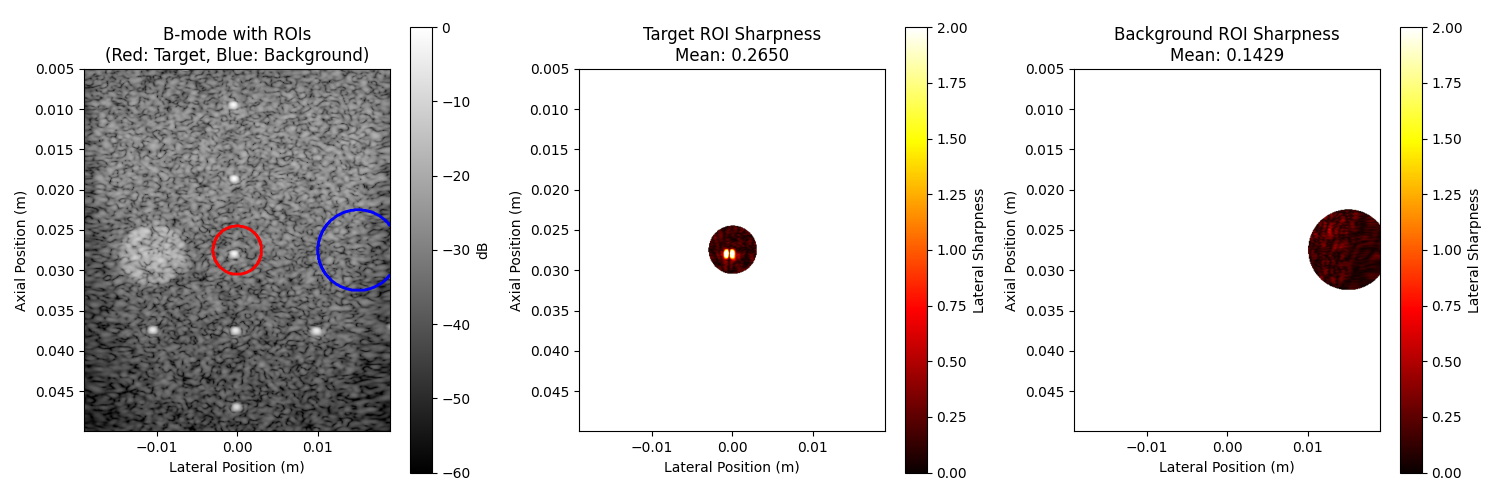
6. Compare sharpness across regions#
We should expect that the point target region is sharper than the background speckle region. Here, we visualize the sharpness in different regions and validate this expectation.
# For the resolution experiment, create ROIs around different areas
center_target = (0.000, 0.0275) # Center of a point target (x, z in meters)
center_background = (0.015, 0.0275) # Background speckle region
# Create circular ROIs
target_radius = 0.003 # 3mm radius
background_radius = 0.005 # 5mm radius
# Build masks for the resolution experiment
mask_target = build_mask(
position=center_target,
dimension=target_radius,
x_axis=scan_resolution.x_axis,
z_axis=scan_resolution.z_axis,
shape="circle",
)
mask_background = build_mask(
position=center_background,
dimension=background_radius,
x_axis=scan_resolution.x_axis,
z_axis=scan_resolution.z_axis,
shape="circle",
)
fig, axes = plt.subplots(1, 3, figsize=(15, 5))
# Show original image with ROI overlays
extent = [x_coords_res.min(), x_coords_res.max(), z_coords_res.max(), z_coords_res.min()]
# Original image
im1 = axes[0].imshow(db_zero(image_resolution), cmap="gray", vmin=vmin, vmax=vmax, extent=extent)
axes[0].contour(
np.linspace(extent[0], extent[1], mask_target.shape[1]),
np.linspace(extent[3], extent[2], mask_target.shape[0]),
mask_target,
colors="red",
linewidths=2,
)
axes[0].contour(
np.linspace(extent[0], extent[1], mask_background.shape[1]),
np.linspace(extent[3], extent[2], mask_background.shape[0]),
mask_background,
colors="blue",
linewidths=2,
)
axes[0].set_title("B-mode with ROIs\n(Red: Target, Blue: Background)")
axes[0].set_xlabel("Lateral Position (m)")
axes[0].set_ylabel("Axial Position (m)")
# Calculate sharpness in different ROIs
sharpness_target = tenengrad(image_resolution, roi=mask_target, normalize=True)
sharpness_background = tenengrad(image_resolution, roi=mask_background, normalize=True)
# Target ROI
masked_target = np.ma.masked_where(~mask_target, sharpness_img_resolution)
im2 = axes[1].imshow(masked_target, cmap="hot", extent=extent, vmin=sharpness_vmin, vmax=sharpness_vmax)
axes[1].set_title(f"Target ROI Sharpness\nMean: {float(sharpness_target):.4f}")
axes[1].set_xlabel("Lateral Position (m)")
axes[1].set_ylabel("Axial Position (m)")
# Background ROI
masked_background = np.ma.masked_where(~mask_background, sharpness_img_resolution)
im3 = axes[2].imshow(masked_background, cmap="hot", extent=extent, vmin=sharpness_vmin, vmax=sharpness_vmax)
axes[2].set_title(f"Background ROI Sharpness\nMean: {float(sharpness_background):.4f}")
axes[2].set_xlabel("Lateral Position (m)")
axes[2].set_ylabel("Axial Position (m)")
# Add colorbars
plt.colorbar(im1, ax=axes[0], label="dB")
plt.colorbar(im2, ax=axes[1], label="Lateral Sharpness")
plt.colorbar(im3, ax=axes[2], label="Lateral Sharpness")
plt.tight_layout()
plt.show()
7. Compare sharpness across regions#
We should expect that the point target region is sharper than the background speckle region.
print("\n=== ROI-based Sharpness Analysis ===")
print(f"Point target region sharpness: {float(sharpness_target):.4f}")
print(f"Background speckle sharpness: {float(sharpness_background):.4f}")
target_bg_ratio = float(sharpness_target) / float(sharpness_background)
print(f"Target/Background ratio: {target_bg_ratio:.2f}")
if target_bg_ratio > 1.5:
print("✅ Point target region is significantly sharper than background!")
print(" This demonstrates the metric's sensitivity to small structures.")
=== ROI-based Sharpness Analysis ===
Point target region sharpness: 0.2650
Background speckle sharpness: 0.1429
Target/Background ratio: 1.85
✅ Point target region is significantly sharper than background!
This demonstrates the metric's sensitivity to small structures.
8. Summary#
We have shown that the Tenengrad sharpness metric can be used to compare the sharpness of different regions in an image.
This metric can be valuable for [2]:
Assessing ultrasound image focusing quality
Comparing different imaging parameters or techniques
Automated quality control in ultrasound systems
Total running time of the script: (0 minutes 1.464 seconds)