Note
Go to the end to download the full example code.
Loading and Visualizing Ultrasound Images#
This example demonstrates how to:
List available datasets
Download a PICMUS dataset using the public API
Visualize the beamformed ultrasound data
This serves as a minimal example of using the ultrasound_metrics.data module. The PICMUS (Plane-wave Imaging Challenge in Medical UltraSound) dataset [1] provides standardized ultrasound data for algorithm comparison.
References#
Example#
1. Import required modules and functions#
import matplotlib.pyplot as plt
from ultrasound_metrics.data import (
create_bmode_image,
db_zero,
list_available_datasets,
)
from ultrasound_metrics.data.uff import (
load_uff_dataset,
)
2. List available datasets#
Available datasets:
- picmus_resolution_experiment: PICMUS challenge resolution/distortion test (experiment)
- picmus_contrast_experiment: PICMUS challenge contrast/speckle test (experiment)
3. Choose a dataset to visualize#
dataset_name = "picmus_resolution_experiment"
print(f"Loading dataset: {dataset_name}")
Loading dataset: picmus_resolution_experiment
4. Load the dataset#
beamformed_image, scan = load_uff_dataset(dataset_name)
print("Dataset loaded successfully!")
print(f"Data shape: {beamformed_image.shape}")
print(f"Data type: {beamformed_image.dtype}")
print(f"Data range: [{beamformed_image.min():.3f}, {beamformed_image.max():.3f}]")
print(f"Scan geometry: x={scan.x_axis.size}, z={scan.z_axis.size}")
Dataset loaded successfully!
Data shape: (387, 609)
Data type: complex64
Data range: [-56.675+21.870j, 52.407-5.226j]
Scan geometry: x=387, z=609
5. Convert to B-mode magnitude for visualization#
data_array = create_bmode_image(beamformed_image)
print(f"After taking magnitude - Data range: [{data_array.min():.3f}, {data_array.max():.3f}]")
After taking magnitude - Data range: [0.003, 62.383]
6. Simple visualization with physical coordinates#
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(12, 5))
# Get physical coordinates from scan geometry
x_coords = scan.x_axis
z_coords = scan.z_axis
# Plot the magnitude data with physical coordinates
# Use 'equal' aspect for square pixels, assuming isotropic grid spacing
im1 = ax1.imshow(
data_array.T,
cmap="gray",
aspect="equal",
extent=[x_coords.min(), x_coords.max(), z_coords.max(), z_coords.min()],
)
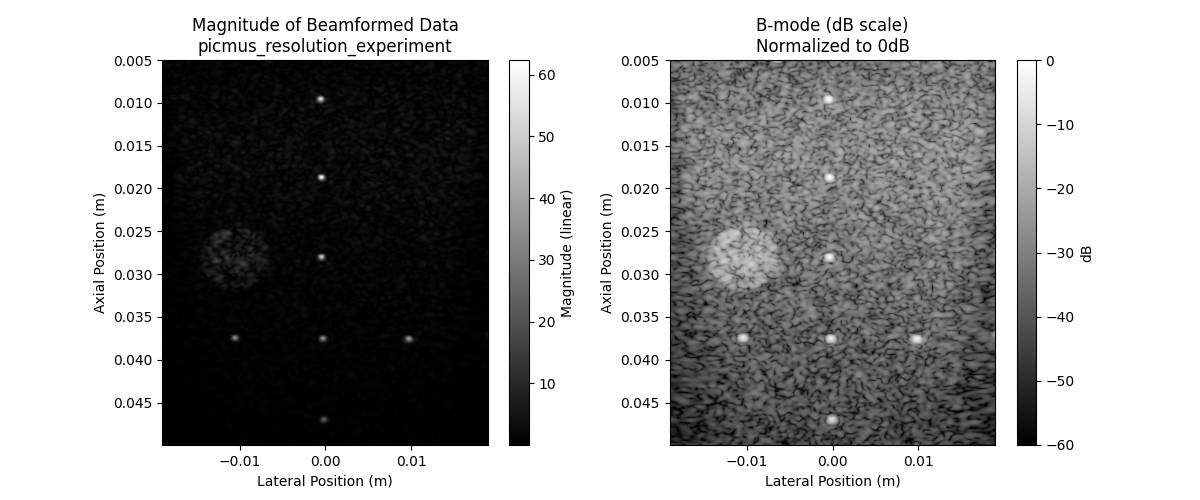
ax1.set_title(f"Magnitude of Beamformed Data\n{dataset_name}")
ax1.set_xlabel("Lateral Position (m)")
ax1.set_ylabel("Axial Position (m)")
# Add colorbar for linear magnitude image
fig.colorbar(im1, ax=ax1, orientation="vertical", label="Magnitude (linear)")
# Plot in dB scale (common for ultrasound visualization)
data_db = db_zero(data_array)
im2 = ax2.imshow(
data_db.T,
cmap="gray",
aspect="equal",
vmin=-60,
vmax=0,
extent=[x_coords.min(), x_coords.max(), z_coords.max(), z_coords.min()],
)
ax2.set_title("B-mode (dB scale)\nNormalized to 0dB")
ax2.set_xlabel("Lateral Position (m)")
ax2.set_ylabel("Axial Position (m)")
# Add colorbar for dB scale image
fig.colorbar(im2, ax=ax2, orientation="vertical", label="dB")
<matplotlib.colorbar.Colorbar object at 0x7f672ea007d0>
print("\nVisualization complete!")
print("Left: Raw beamformed data")
print("Right: B-mode in dB scale (typical ultrasound display)")
print("\nNote: This example demonstrates the new module structure:")
print("- General utilities (create_bmode_image, db_zero) from ultrasound_metrics.data")
print("- UFF-specific functionality (load_uff_dataset) from ultrasound_metrics.data.uff")
Visualization complete!
Left: Raw beamformed data
Right: B-mode in dB scale (typical ultrasound display)
Note: This example demonstrates the new module structure:
- General utilities (create_bmode_image, db_zero) from ultrasound_metrics.data
- UFF-specific functionality (load_uff_dataset) from ultrasound_metrics.data.uff
Total running time of the script: (0 minutes 0.192 seconds)