Note
Go to the end to download the full example code.
Generalized Contrast-to-Noise Ratio (gCNR) Metric#
This example demonstrates how to:
Load an ultrasound dataset
Choose ROI selection mode (interactive or hardcoded)
Visualize the B-mode image and select signal/noise ROIs
Print ROI summary and statistics
Compute and print the gCNR (optionally plot PDFs)
This workflow is backend-agnostic and uses the helper functions from ultrasound_metrics. The gCNR metric follows the formulation in [1] and uses the PICMUS dataset [2].
References#
Example#
1. Import required modules and functions#
import matplotlib.pyplot as plt
import numpy as np
from ultrasound_metrics.data import db_zero
# (Optional) for interactive ROI selection
from ultrasound_metrics.interactive.napari_utils import load_ultrasound_for_napari, select_rois_with_labels
from ultrasound_metrics.metrics.gcnr import gcnr
from ultrasound_metrics.roi.masks import build_mask
2. Set ROI selection mode#
def is_headless_environment():
"""Detect if running in headless environment (no GUI available)."""
import os
import sys
# Check for ReadTheDocs
if os.environ.get("READTHEDOCS") == "True":
return True
# Check for sphinx-gallery module
return "sphinx_gallery" in sys.modules
# Interactive if not headless (feel free to change to True / False for local testing)
use_interactive = not is_headless_environment()
3. Load the ultrasound dataset#
image_data, metadata, scan = load_ultrasound_for_napari("picmus_resolution_experiment")
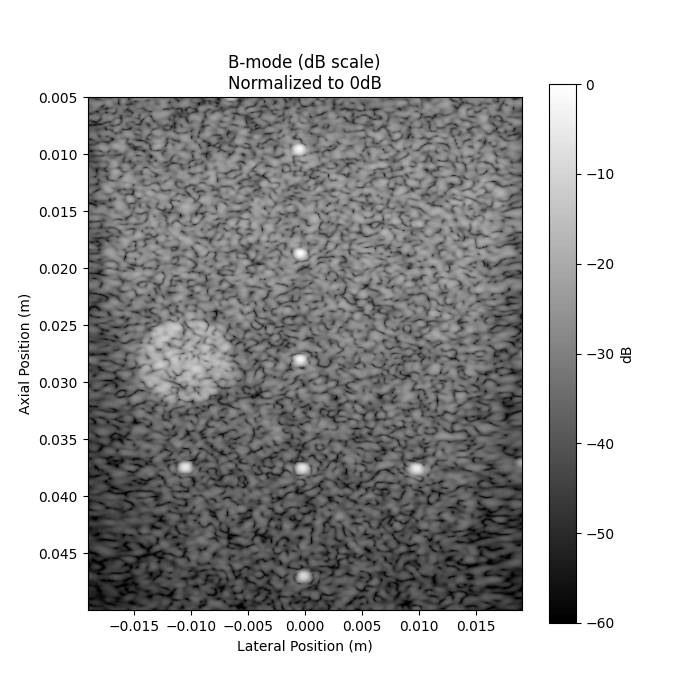
4. Visualize the B-mode image to help select ROI center and radii#
x_coords = metadata["x_axis"]
z_coords = metadata["z_axis"]
# Only convert to dB if not already in dB scale
if metadata.get("use_db_scale", True):
image_db = image_data
else:
image_db = db_zero(image_data)
if not use_interactive:
plt.figure(figsize=(7, 7))
plt.imshow(
image_db,
cmap="gray",
aspect="equal",
vmin=-60,
vmax=0,
extent=[x_coords.min(), x_coords.max(), z_coords.max(), z_coords.min()],
)
plt.title("B-mode (dB scale)\nNormalized to 0dB")
plt.xlabel("Lateral Position (m)")
plt.ylabel("Axial Position (m)")
plt.colorbar(label="dB")
plt.show()

5. Select ROIs (either interactively or with hardcoded masks)#
if use_interactive:
roi_result = select_rois_with_labels(image=image_data)
mask_signal = roi_result["mask_signal"]
mask_noise = roi_result["mask_noise"]
else:
# Hardcoded ROI selection and visualization
center = (-0.0105, 0.028) # (x, z) in meters
signal_radius = 0.004 # 4 mm
noise_radius = 0.0075 # 7.5 mm
mask_signal = build_mask(
position=center, dimension=signal_radius, x_axis=metadata["x_axis"], z_axis=metadata["z_axis"], shape="circle"
)
mask_noise_outer = build_mask(
position=center, dimension=noise_radius, x_axis=metadata["x_axis"], z_axis=metadata["z_axis"], shape="circle"
)
mask_noise = np.logical_and(mask_noise_outer, ~mask_signal)
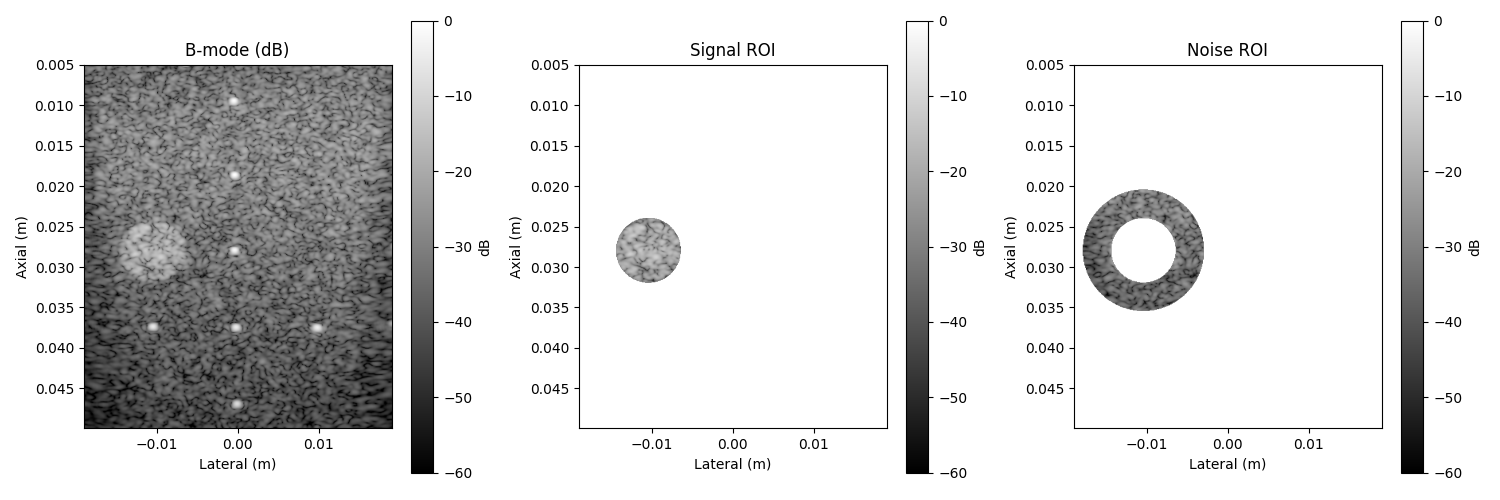
# --- Visualization of B-mode, signal, and noise masks ---
fig, axs = plt.subplots(1, 3, figsize=(15, 5))
bmode_args = {
"extent": [x_coords.min(), x_coords.max(), z_coords.max(), z_coords.min()],
"cmap": "gray",
"vmin": -60,
"vmax": 0,
}
im0 = axs[0].imshow(image_db, **bmode_args)
axs[0].set_title("B-mode (dB)")
axs[0].set_xlabel("Lateral (m)")
axs[0].set_ylabel("Axial (m)")
fig.colorbar(im0, ax=axs[0], orientation="vertical", label="dB")
masked_signal = np.ma.masked_where(~mask_signal, image_db)
im1 = axs[1].imshow(masked_signal, **bmode_args)
axs[1].set_title("Signal ROI")
axs[1].set_xlabel("Lateral (m)")
axs[1].set_ylabel("Axial (m)")
fig.colorbar(im1, ax=axs[1], orientation="vertical", label="dB")
masked_noise = np.ma.masked_where(~mask_noise, image_db)
im2 = axs[2].imshow(masked_noise, **bmode_args)
axs[2].set_title("Noise ROI")
axs[2].set_xlabel("Lateral (m)")
axs[2].set_ylabel("Axial (m)")
fig.colorbar(im2, ax=axs[2], orientation="vertical", label="dB")
plt.tight_layout()
plt.show()
6. Print ROI summary#
signal_pixels = mask_signal.sum()
noise_pixels = mask_noise.sum()
print("\n=== ROI Selection Summary ===")
print(f"Signal region pixel count: {signal_pixels}")
print(f"Noise region pixel count: {noise_pixels}")
=== ROI Selection Summary ===
Signal region pixel count: 6895
Noise region pixel count: 17361
7. Print basic statistics#
values_signal = image_data[mask_signal]
values_noise = image_data[mask_noise]
print("\n=== Region Statistics ===")
print(f"Signal mean: {values_signal.mean():.3f} dB")
print(f"Noise mean: {values_noise.mean():.3f} dB")
=== Region Statistics ===
Signal mean: -23.187 dB
Noise mean: -35.079 dB
8. Compute and print gCNR (optionally plot PDFs)#
fig, ax = plt.subplots(figsize=(6, 4))
gcnr_value = gcnr(values_inside=values_signal, values_outside=values_noise, ax=ax)
plt.tight_layout()
plt.show()
print("\n=== gCNR Results ===")
print(f"gCNR: {gcnr_value:.3f}")
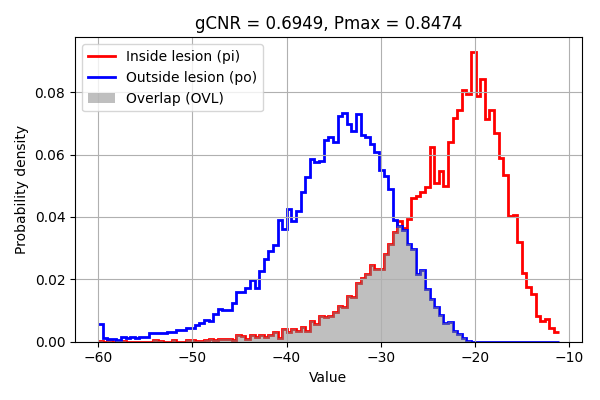
=== gCNR Results ===
gCNR: 0.695
Total running time of the script: (0 minutes 0.572 seconds)