Note
Go to the end to download the full example code.
Signal-to-Noise Ratio (SNR) for Speckle Regions#
This example demonstrates the Signal-to-Noise Ratio (SNR) metric for ultrasound speckle analysis. We explore how SNR varies in different tissue regions and validate measurements against theoretical Rayleigh distribution values.
Mathematical Definition#
The Signal-to-Noise Ratio for ultrasound speckle is defined as the ratio of mean to standard deviation:
where \(\mathbb{E}[X]\) is the mean and \(\sqrt{\text{Var}(X)}\) is the standard deviation of the envelope-detected signal values.
Fully developed speckle follows a Rayleigh distribution [1], so the theoretical SNR is:
This metric provides insight into speckle quality and can be used to:
Assess fully developed speckle regions
Compare different beamforming techniques
Validate speckle statistics in phantom studies
Quality control in ultrasound systems
ROI Selection Best Practices:
Following Perdios et al. [2], speckle patterns should be assessed within regions of approximately 10λx10λ (where λ is the wavelength) to ensure sufficient statistical sampling while maintaining spatial homogeneity. Regions should be placed away from boundaries and avoid compounding artifacts near field edges.
References#
Example#
Import required modules and functions#
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from matplotlib.patches import Circle
from ultrasound_metrics.data import db_zero
from ultrasound_metrics.data.uff import load_uff_dataset
from ultrasound_metrics.metrics.snr import snr
from ultrasound_metrics.roi.masks import build_mask
Load PICMUS datasets [3] for comparison#
# Load contrast/speckle dataset (contains fully developed speckle regions)
image_contrast, scan_contrast = load_uff_dataset("picmus_contrast_experiment")
image_contrast = image_contrast.T
# Load resolution dataset (contains point targets and speckle)
image_resolution, scan_resolution = load_uff_dataset("picmus_resolution_experiment")
image_resolution = image_resolution.T
Visualize datasets side by side#
fig, axes = plt.subplots(1, 2, figsize=(12, 6))
# Contrast/speckle dataset
extent_contrast = [
scan_contrast.x_axis.min(),
scan_contrast.x_axis.max(),
scan_contrast.z_axis.max(),
scan_contrast.z_axis.min(),
]
axes[0].imshow(db_zero(image_contrast), cmap="gray", extent=extent_contrast, vmin=-60, vmax=0)
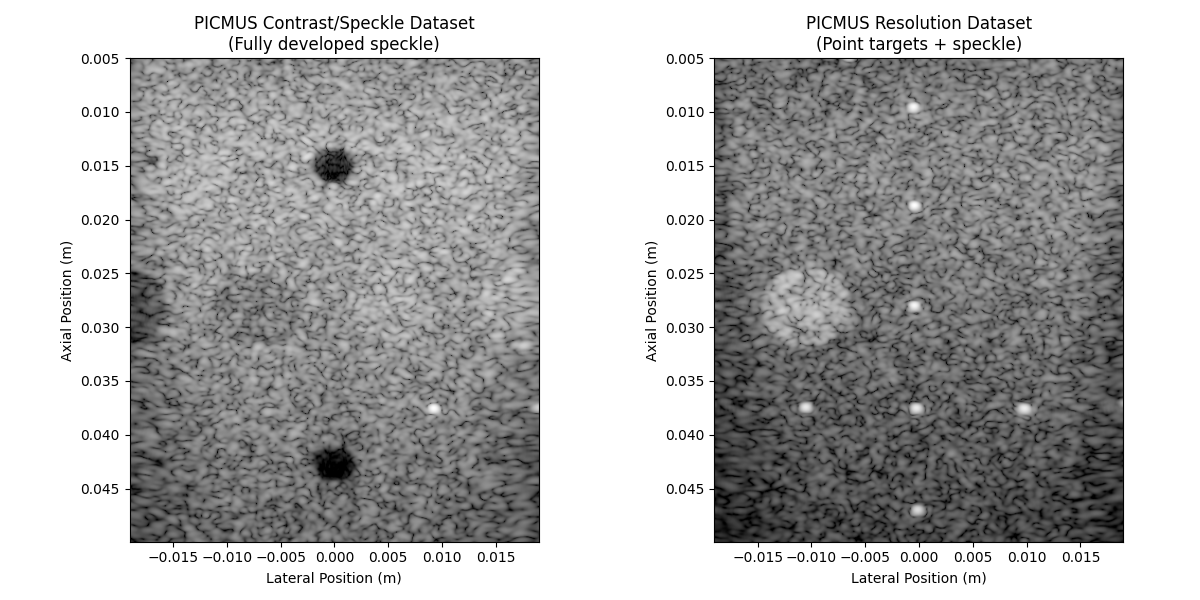
axes[0].set_title("PICMUS Contrast/Speckle Dataset\n(Fully developed speckle)")
axes[0].set_xlabel("Lateral Position (m)")
axes[0].set_ylabel("Axial Position (m)")
# Resolution dataset
extent_resolution = [
scan_resolution.x_axis.min(),
scan_resolution.x_axis.max(),
scan_resolution.z_axis.max(),
scan_resolution.z_axis.min(),
]
axes[1].imshow(db_zero(image_resolution), cmap="gray", extent=extent_resolution, vmin=-60, vmax=0)
axes[1].set_title("PICMUS Resolution Dataset\n(Point targets + speckle)")
axes[1].set_xlabel("Lateral Position (m)")
axes[1].set_ylabel("Axial Position (m)")
plt.tight_layout()

Define ROI locations for speckle analysis#
# Following Perdios et al., we use regions of approximately 10λx10λ
# Assuming center frequency ~5 MHz and speed of sound ~1540 m/s
# λ ≈ 1540/5e6 ≈ 0.3 mm, so 10λ ≈ 3 mm radius
roi_radius = 0.003 # 3 mm radius
# Equation S9 from Perdios et al. [2]_ and Equation 4 from Burckhardt [1]_
theoretical_rayleigh_snr = np.sqrt(np.pi / (4 - np.pi))
# Define ROI locations for different types of regions
roi_specs = [
{
"name": "Fully Developed Speckle (contrast dataset)",
"dataset": "contrast",
"center": (-0.005, 0.035),
"expected_snr": theoretical_rayleigh_snr,
"description": "Homogeneous speckle region",
},
{
"name": "Fully Developed Speckle (resolution dataset)",
"dataset": "resolution",
"center": (0.008, 0.030),
"expected_snr": theoretical_rayleigh_snr,
"description": "Homogeneous speckle region",
},
{
"name": "Point Target Region",
"dataset": "resolution",
"center": (0.000, 0.0275), # Near a point target
"expected_snr": None, # Should deviate from Rayleigh
"description": "Point target + speckle",
},
{
"name": "Lesion Boundary",
"dataset": "contrast",
"center": (-0.0025, 0.028), # Near lesion boundary
"expected_snr": None, # Mixed statistics
"description": "Lesion boundary region",
},
]
Compute SNR for each ROI#
results_data = []
for spec in roi_specs:
# Select the appropriate dataset and scan
if spec["dataset"] == "contrast":
image = image_contrast
scan = scan_contrast
extent = extent_contrast
else:
image = image_resolution
scan = scan_resolution
extent = extent_resolution
# Create ROI mask
mask = build_mask(
position=spec["center"],
dimension=roi_radius,
x_axis=scan.x_axis,
z_axis=scan.z_axis,
shape="circle",
)
# Extract values in ROI
roi_values = image[mask]
# Compute SNR (function handles complex data internally)
snr_linear = snr(roi_values, db=False)
snr_db = snr(roi_values, db=True)
# Calculate deviation from theoretical Rayleigh SNR if expected
if spec["expected_snr"] is not None:
deviation = abs(snr_linear - spec["expected_snr"])
deviation_percent = (deviation / spec["expected_snr"]) * 100
else:
deviation = None
deviation_percent = None
results_data.append({
"ROI": spec["name"],
"Dataset": spec["dataset"],
"SNR (linear)": snr_linear,
"SNR (dB)": snr_db,
"Expected SNR": spec["expected_snr"],
"Deviation": deviation,
"Deviation (%)": deviation_percent,
"Description": spec["description"],
"mask": mask,
"center": spec["center"],
"extent": extent,
"image": image,
})
Display results table#
results_df = pd.DataFrame(results_data)
print("=== SNR Analysis Results ===")
display_cols = ["ROI", "Dataset", "SNR (linear)", "SNR (dB)", "Expected SNR", "Deviation (%)"]
print(results_df[display_cols].round(3))
print(f"\nTheoretical Rayleigh SNR: {theoretical_rayleigh_snr:.3f}")
for _, row in results_df.iterrows():
if row["Expected SNR"] is not None:
if row["Deviation (%)"] < 10:
status = "✅ Good agreement with theory (Rayleigh distribution)"
elif row["Deviation (%)"] < 20:
status = "⚠️ Moderate deviation from theory"
else:
status = "❌ Significant deviation from theory"
print(f"{row['ROI']}: SNR = {row['SNR (linear)']:.3f}, {status}")
else:
print(f"{row['ROI']}: SNR = {row['SNR (linear)']:.3f} (non-Rayleigh distribution)")
=== SNR Analysis Results ===
ROI ... Deviation (%)
0 Fully Developed Speckle (contrast dataset) ... 2.189
1 Fully Developed Speckle (resolution dataset) ... 1.115
2 Point Target Region ... NaN
3 Lesion Boundary ... NaN
[4 rows x 6 columns]
Theoretical Rayleigh SNR: 1.913
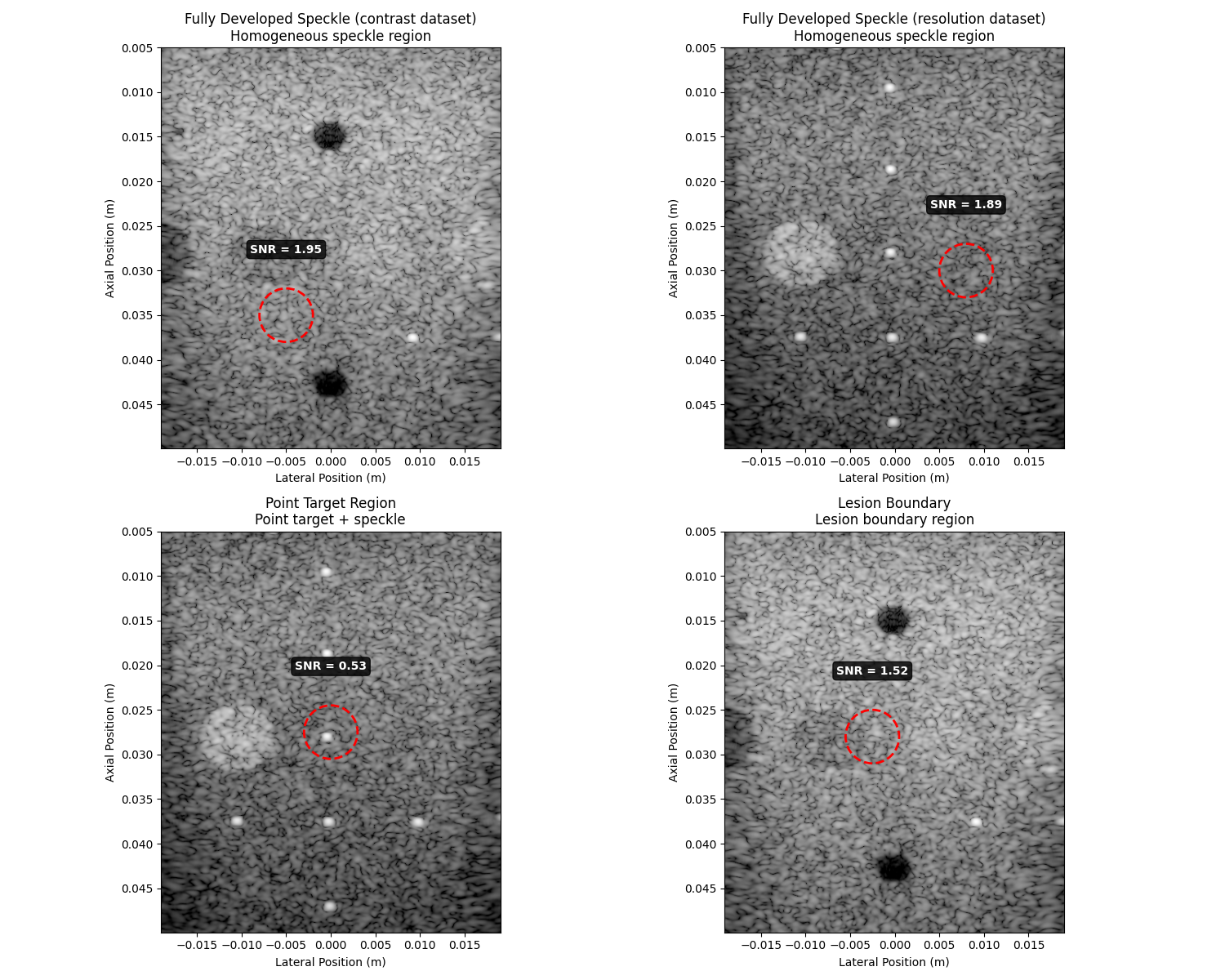
Fully Developed Speckle (contrast dataset): SNR = 1.955, ✅ Good agreement with theory (Rayleigh distribution)
Fully Developed Speckle (resolution dataset): SNR = 1.892, ✅ Good agreement with theory (Rayleigh distribution)
Point Target Region: SNR = 0.534, ❌ Significant deviation from theory
Lesion Boundary: SNR = 1.520, ❌ Significant deviation from theory
Visualize ROIs and SNR measurements#
fig, axes = plt.subplots(2, 2, figsize=(15, 12))
axes = axes.flatten()
for i, (_, row) in enumerate(results_df.iterrows()):
ax = axes[i]
# Display the image
ax.imshow(db_zero(row["image"]), cmap="gray", extent=row["extent"], vmin=-60, vmax=0)
# Overlay ROI circle
circle = Circle(row["center"], roi_radius, fill=False, color="red", linewidth=2, linestyle="--")
ax.add_patch(circle)
# Add SNR annotation
ax.text(
row["center"][0],
row["center"][1] - 0.008,
f"SNR = {row['SNR (linear)']:.2f}",
ha="center",
va="top",
color="white",
fontweight="bold",
bbox={"boxstyle": "round,pad=0.3", "facecolor": "black", "alpha": 0.8},
)
ax.set_title(f"{row['ROI']}\n{row['Description']}")
ax.set_xlabel("Lateral Position (m)")
ax.set_ylabel("Axial Position (m)")
plt.tight_layout()

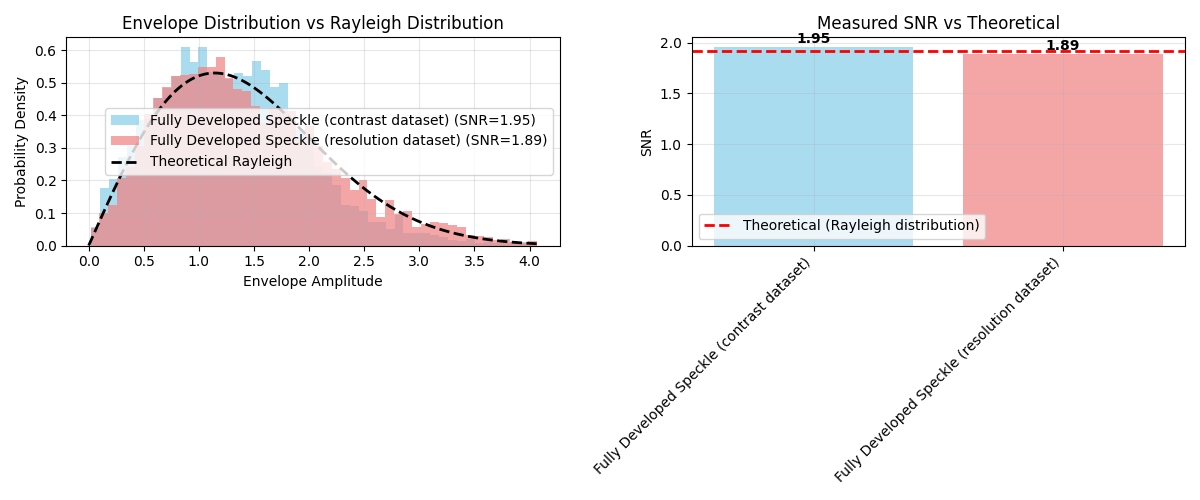
Statistical analysis of speckle regions#
# Focus on the fully developed speckle regions
speckle_rows = results_df[results_df["Expected SNR"].notna()]
fig, axes = plt.subplots(1, 2, figsize=(12, 5))
colors = ["skyblue", "lightcoral"]
for i, (_, row) in enumerate(speckle_rows.iterrows()):
# Extract ROI values and convert to envelope (magnitude)
roi_values_complex = row["image"][row["mask"]]
roi_values = np.abs(roi_values_complex) # Take envelope for histogram
# Create histogram of envelope values
axes[0].hist(
roi_values,
bins=50,
alpha=0.7,
density=True,
color=colors[i],
label=f"{row['ROI']} (SNR={row['SNR (linear)']:.2f})",
)
# Overlay theoretical Rayleigh distribution
x_theory = np.linspace(0, roi_values.max(), 1000)
# For comparison, use sigma that gives the same mean as our data
sigma_rayleigh = np.mean(roi_values) / np.sqrt(np.pi / 2)
rayleigh_pdf = (x_theory / sigma_rayleigh**2) * np.exp(-(x_theory**2) / (2 * sigma_rayleigh**2))
axes[0].plot(x_theory, rayleigh_pdf, "k--", linewidth=2, label="Theoretical Rayleigh")
axes[0].set_xlabel("Envelope Amplitude")
axes[0].set_ylabel("Probability Density")
axes[0].set_title("Envelope Distribution vs Rayleigh Distribution")
axes[0].legend()
axes[0].grid(True, alpha=0.3)
# SNR comparison bar chart
roi_names = [row["ROI"] for _, row in speckle_rows.iterrows()]
snr_values = [row["SNR (linear)"] for _, row in speckle_rows.iterrows()]
bars = axes[1].bar(roi_names, snr_values, alpha=0.7, color=colors)
axes[1].axhline(
y=theoretical_rayleigh_snr,
color="red",
linestyle="--",
linewidth=2,
label="Theoretical (Rayleigh distribution)",
)
axes[1].set_ylabel("SNR")
axes[1].set_title("Measured SNR vs Theoretical")
axes[1].legend()
axes[1].grid(True, alpha=0.3)
# Add value labels on bars
for bar, value in zip(bars, snr_values):
height = bar.get_height()
axes[1].text(
bar.get_x() + bar.get_width() / 2.0,
height + 0.01,
f"{value:.2f}",
ha="center",
va="bottom",
fontweight="bold",
)
plt.xticks(rotation=45, ha="right")
plt.tight_layout()
%% Show all figures (if run as a script)
plt.show()
Best Practices for SNR Measurement#
Based on the analysis above, here are key guidelines for SNR measurement:
ROI Size Guidelines:
Use regions of approximately 10λx10λ for adequate statistical sampling
For 5 MHz transducers: ≈3mm x 3mm ROI provides good sampling
Larger ROIs provide more stable statistics but may include spatial variations
Region Selection Criteria:
Choose homogeneous speckle areas away from boundaries
Avoid edges with incomplete plane wave coverage
Regions to Avoid:
Lesion edges (mixed Rayleigh/non-Rayleigh statistics)
Point targets (coherent scattering, non-Rayleigh)
Field edges (incomplete acoustic data)
Areas with obvious artifacts
Expected Values for Quality Assessment:
Clinical and Research Applications:
Speckle quality assessment in different tissue types
Beamforming algorithm evaluation and optimization
System validation and quality control
Total running time of the script: (0 minutes 1.103 seconds)